One of the greatest mysteries in our Universe is right here on our own doorstep. No, closer - it's in every fibre of our being.
At least 3.7 billion years ago, a few simple molecules worked together to create something new. Then a few more. And, somehow, these snowballing combinations eventually produced the first very basic living organisms that would evolve and branch out to become all life on Earth.
We don't know the order it happened in; heck, we don't even know when or where it happened. But new research is showing us the possibilities.
With a specially developed software tool, scientists in Poland and South Korea have traced the potential synthesis routes from the simple precursor molecules (prebiotics) present on early Earth to the complex biomolecules that gave rise to the planet's teeming life today.
"Although hundreds of organic reactions have been validated under consensus prebiotic conditions, we still have only a fragmentary understanding of how these individual steps combined into complete synthetic pathways to generate life's building blocks," the researchers wrote in their paper.
"We harnessed the power of computer-assisted organic synthesis to map the network of molecules that are synthesisable from basic prebiotic feedstocks."
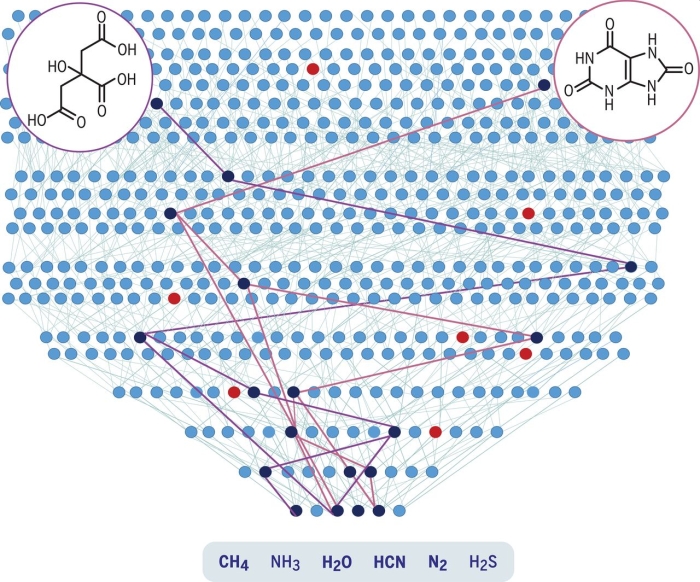 Illustration of the reaction tree. (Wołos et al., Science, 2020)
Illustration of the reaction tree. (Wołos et al., Science, 2020)
The software is called Allchemy, and the team developed it by encoding known prebiotic chemical reactions in a format that a machine can read. They included information about incompatible groups and conditions, and known reaction mechanisms under prebiotic conditions.
Six basic known prebiotic molecules that existed before life on Earth are fed into the program as a starting point - water, nitrogen, cyanide, ammonia, methane, and sulfur.
From there, the algorithm identifies possible successive reactions. These start with the original six molecules, and then iteratively progress, growing more and more complex. Each newly created molecule coan react with the products of all previous iterations in the chain, as well as the original six molecules.
Eventually, these chain reactions produce the building blocks of life, such as nucleic acids, lipids, and proteins like enzymes.
The team ran their software for several generations, and reproduced all the known pathways to prebiotic molecules.
Some prebiotic molecules, Allchemy identified, can form incredibly quickly - glycine, which our bodies use as a neurotransmitter, forms after just one reaction.
Urea, adenine (a component of DNA), butenedioic acid, and oxalic acid can form after just two reactions; and glyceraldehyde, isoguanine, aspartic acid, hypoxanthine, cytosine, phenylalanine, succinic acid, malic acid, glyoxylic acid and aldotetrose can form after three.
Within two hours, the software had produced 82 biotic molecules and 36,603 abiotic molecules, the researchers said.
In addition to the known pathways, the software identified several previously unknown synthesis routes to prebiotic molecules. These, the team tested by creating the molecules in a laboratory setting, according to the Allchemy recipe.
That would be an interesting enough finding on its own, but Allchemy was far from done. From the tree of life the software generated, the team identified three kinds of chemical emergence that are crucial to life.
Firstly, molecules created within the network can enable new kinds of chemical reactions, vastly expanding the number of potential products from a generation of molecules. Secondly, within a few generations, simple chemical systems emerge - including self-regenerating cycles. These were also verified in lab experiments.
And, finally, the network identified routes to surfactants. These molecules that spontaneously connect with each other to form enclosed bubbles. They form cell walls; without them, we wouldn't have biological compartmentalisation as we know it, which is of crucial importance to the organisation of biomolecular systems.
The actual chemical pathways life took billions of years ago will likely never be known to us. But Allchemy shows that maybe it wasn't as difficult as we might think.
And it could prove to be an extremely useful tool for exploring conditions that could give rise to life, not just on Earth, but possibly elsewhere.
"Taken together, the above analyses and synthetic examples lead us to suggest that computational reaction network algorithms are useful for identifying new synthetic routes to prebiotically relevant targets and indispensable for the discovery of prebiotic chemical systems that are otherwise challenging to discern," the researchers wrote in their paper.
"Naturally, the prebiotic reaction networks should and will grow as distinct prebiotically plausible transformations are experimentally validated."
Allchemy has been made freely available to the public. And the team's research has been published in Science.
