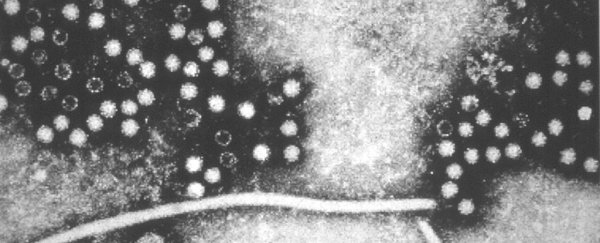
An analysis of the genetic material in the ocean has identified thousands of previously unknown RNA viruses and doubled the number of phyla, or biological groups, of viruses thought to exist, according to a new study our team of researchers has published in the journal Science.
RNA viruses are best known for the diseases they cause in people, ranging from the common cold to COVID-19. They also infect plants and animals important to people.
These viruses carry their genetic information in RNA, rather than DNA. RNA viruses evolve at much quicker rates than DNA viruses do. While scientists have cataloged hundreds of thousands of DNA viruses in their natural ecosystems, RNA viruses have been relatively unstudied.
Unlike humans and other organisms composed of cells, however, viruses lack unique short stretches of DNA that could act as what researchers call a genetic bar code. Without this bar code, trying to distinguish different species of virus in the wild can be challenging.
To get around this limitation, we decided to identify the gene that codes for a particular protein that allows a virus to replicate its genetic material. It is the only protein that all RNA viruses share, because it plays an essential role in how they propagate themselves. Each RNA virus, however, has small differences in the gene that codes for the protein that can help distinguish one type of virus from another.
So we screened a global database of RNA sequences from plankton collected during the four-year Tara Oceans expeditions global research project. Plankton are any aquatic organisms that are small to swim against the current. They're a vital part of ocean food webs and are common hosts for RNA viruses. Our screening ultimately identified over 44,000 genes that code for the virus protein.
Our next challenge, then, was to determine the evolutionary connections between these genes. The more similar two genes were, the more likely viruses with those genes were closely related. Because these sequences had evolved so long ago (possibly predating the first cell), the genetic signposts indicating where new viruses may have split off from a common ancestor had been lost to time.
A form of artificial intelligence called machine learning, however, allowed us to systematically organize these sequences and detect differences more objectively than if the task were done manually.
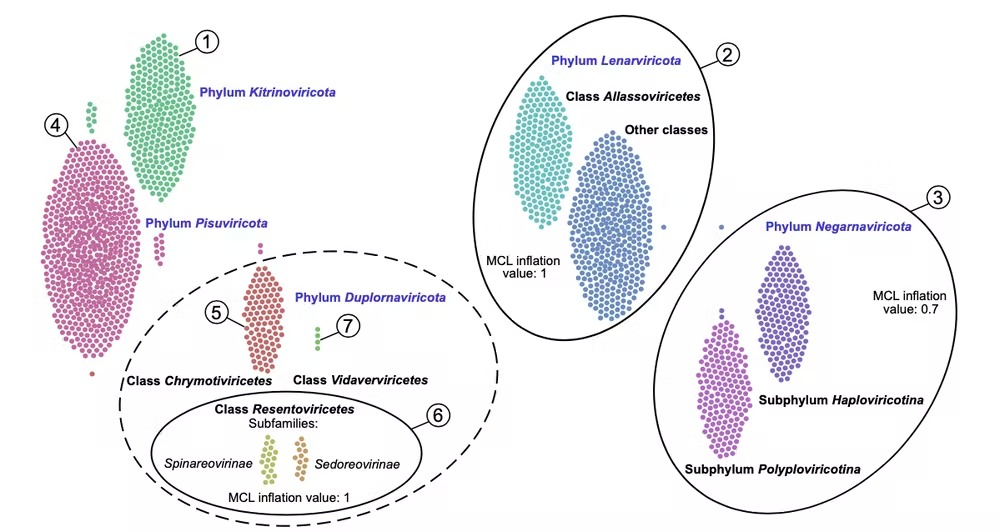 The five existing phyla of RNA viruses organized by the new method. (Zayed et al., Science, 2022)
The five existing phyla of RNA viruses organized by the new method. (Zayed et al., Science, 2022)
We identified a total of 5,504 new marine RNA viruses and doubled the number of known RNA virus phyla from five to 10.
Mapping these new sequences geographically revealed that two of the new phyla were particularly abundant across vast oceanic regions, with regional preferences in either temperate and tropical waters (the Taraviricota, named after the Tara Oceans expeditions) or the Arctic Ocean (the Arctiviricota).
We believe that Taraviricota might be the missing link in the evolution of RNA viruses that researchers have long sought, connecting two different known branches of RNA viruses that diverged in how they replicate.
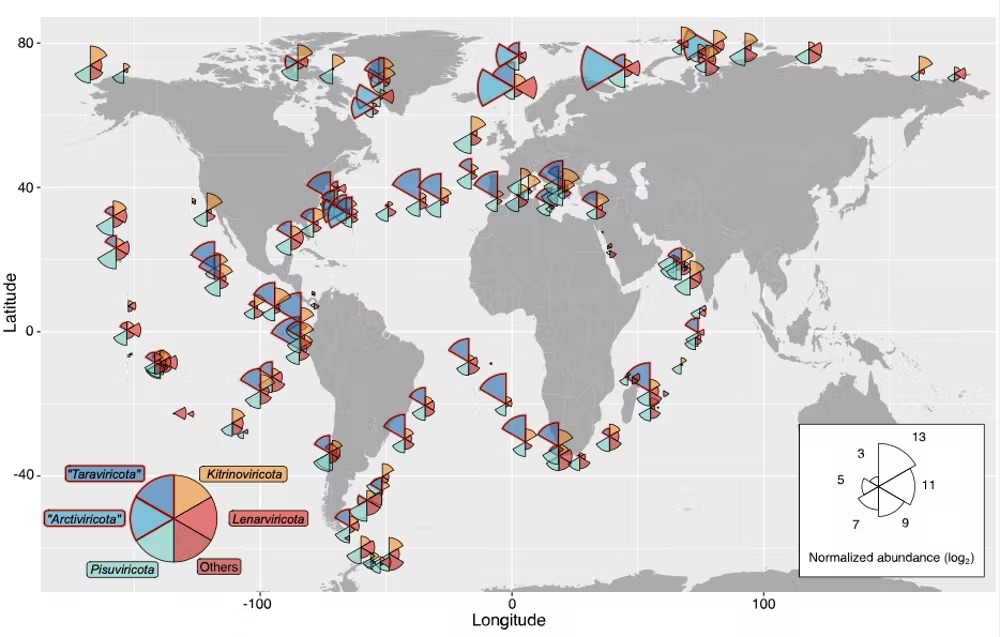 Map showing the distribution of RNA viruses across the ocean. (Zayed et al., Science, 2022)
Map showing the distribution of RNA viruses across the ocean. (Zayed et al., Science, 2022)
Above: Wedge size is proportional to the average abundance of viruses present in that area, and wedge color indicates virus phyla.
Why it matters
These new sequences help scientists better understand not only the evolutionary history of RNA viruses but also the evolution of early life on Earth.
As the COVID-19 pandemic has shown, RNA viruses can cause deadly diseases. But RNA viruses also play a vital role in ecosystems because they can infect a wide array of organisms, including microbes that influence environments and food webs at the chemical level.
Mapping out where in the world these RNA viruses live can help clarify how they affect the organisms driving many of the ecological processes that run our planet. Our study also provides improved tools that can help researchers catalog new viruses as genetic databases grow.

What still isn't known
Despite identifying so many new RNA viruses, it remains challenging to pinpoint what organisms they infect. Researchers are also currently limited to mostly fragments of incomplete RNA virus genomes, partly because of their genetic complexity and technological limitations.
Our next steps would be to figure out what kinds of genes might be missing and how they changed over time. Uncovering these genes could help scientists better understand how these viruses work. ![]()
Guillermo Dominguez Huerta, Science Consultant in Microbiology, The Ohio State University; Ahmed Zayed, Research Scientist in Microbiology, The Ohio State University; James Wainaina, Postdoctoral Research Associate in Microbiology, The Ohio State University, and Matthew Sullivan, Professor of Microbiology, The Ohio State University.
This article is republished from The Conversation under a Creative Commons license. Read the original article.