You may have heard of Folding@home, the number-crunching app you can run on your computer to help researchers tackle certain medical problems, including the new coronavirus. In the past month, the network of volunteers who've installed it has become so vast, the platform is outperforming the most powerful supercomputers in the world.
According to the Folding@home director, biochemist Greg Bowman, some 700,000 new Folding@home operators have joined up in recent weeks. That's a huge increase over the 30,000 people who are typically running Folding@home at any one time.
And it makes a huge difference in computing power too – the Folding@home network reached an astounding 2.4 exaFLOPs of processing power earlier this week, making it faster than the top 500 supercomputers in the world combined.
To give you some kind of idea of scale, the new Xbox One Series X console appearing this year is promising 12 teraFLOPs of graphics processing power, and you need a million teraflops to make up an exaFLOP.
It shows the power of ordinary home computers working at scale, and all this additional performance is being put to good use. The primary job of Folding@home is to model how proteins behave in the body, underpinning so many core biological functions.
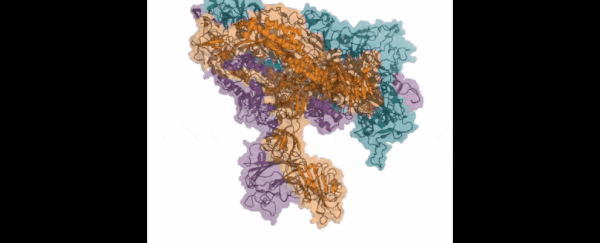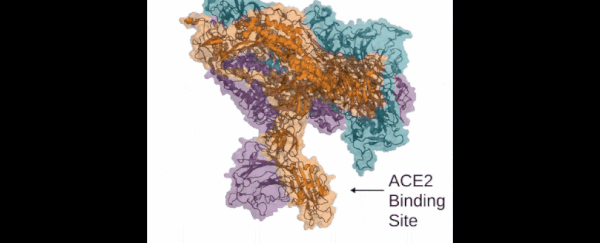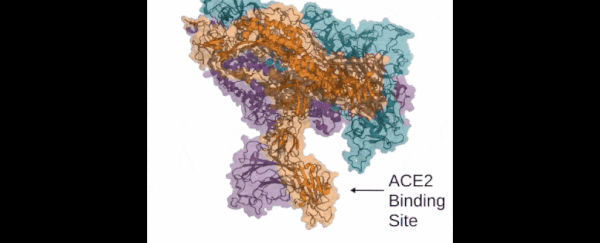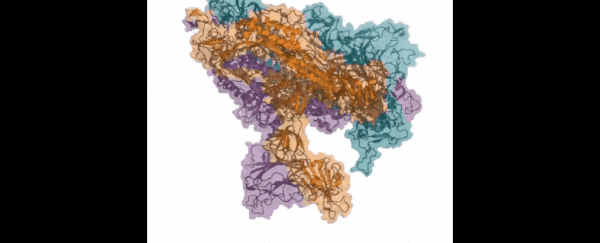
Those functions include virus infection: the Folding@home team is looking in particular at how the so-called 'spike' of the SARS-CoV-2 virus (which is actually made up of three proteins) attaches itself to human cells and infects the human body.
Those three proteins the spike uses to grab hold of the ACE2 human cells look so much like the mouth of the Demogorgon from Stranger Things, the research team has even nicknamed it after the monster.
This is the key way that the new coronavirus can penetrate tissue in the human body, and so blocking it could be crucial to future therapies and treatments. If we can understand more about how the spike proteins work – which is what the computer simulations powered by Folding@home are doing – then we can better design the drugs to stop them.
"If you tried to simulate the opening of the spike on your home computer, you'd be lucky to see even part of the process within the next 100 years," writes Bowman on the Folding@home blog. "Fortunately, we have reinforcements!"
The magic of Folding@home is that it splits up complex protein computer modelling into smaller tasks that can then be distributed to thousands of computers around the world – everyone takes a chunk.
If you install the app, you decide when it runs and how much of your computer's processing power it uses (you don't need a particularly high-end computer at all). If you want, the app will run when it detects you're not using the computer for anything else.
Plenty of other coronavirus-related projects are underway at Folding@home too: data are being processed to assess the effectiveness of potential drugs now being tested in the lab, and to analyse how the coronavirus controls a cell's machinery after infection.
No matter how slow your computer or how little time you think it can spend performing calculations for Folding@home, every effort will get us closer to better treatments for and protections against COVID-19.
Having reached 700,000 folders, Folding@home is now hoping to breach the one million mark. You can download the software for free from here.