Scientists have just discovered an entirely new way that DNA can be synthesized.
The business of constructing DNA (deoxyribonucleic acid, to give it its full name) usually requires a template that builder proteins called enzymes can work from.
But now, a team from Stanford University has found that a type of enzyme known as a polymerase can work without a blueprint. Its shape itself acts as a mold that new DNA can be synthesized from, with no external reference materials required.
This has never been seen before.
While it's not quite a discovery that rewrites the science textbooks, it certainly adds an intriguing new chapter (or three). There are implications for bacteria behavior, biological evolution, and the building blocks of life.
"The enzymatic synthesis of nucleic acids is a fundamental process that underlies genome replication, repair, and diverse forms of information processing across all domains of life," write the researchers in their published paper.
"These findings expand the functional landscape of nucleic acid polymerases, revealing a protein-templated mechanism for sequence-specific DNA synthesis."
The impetus behind the study was to look at defense-associated reverse transcriptases (DRTs), which bacteria use to fend off attacks by viruses. Scientists have already seen some unusual DNA-building behavior in these polymerases.
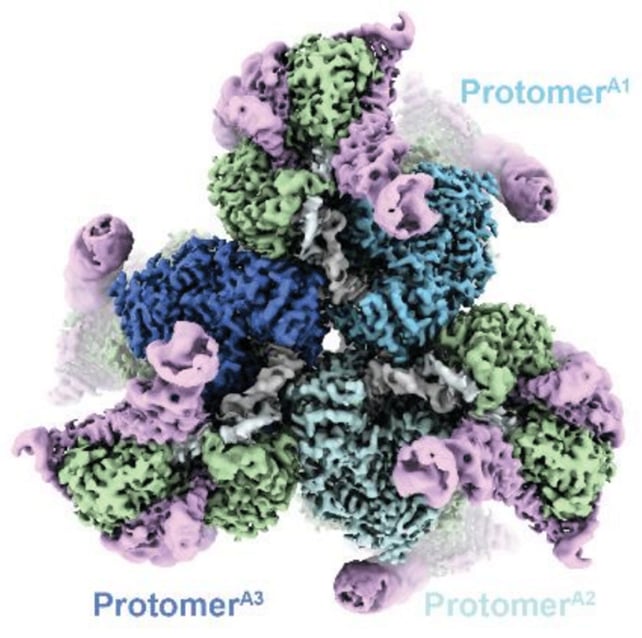
Specifically, the team cloned a DRT3 system from Escherichia coli bacteria, examining its behavior in test tubes and in living cells. That revealed three parts of a DNA-making machine: two enzymes called Drt3a and Drt3b, plus a piece of non-coding RNA.
It was Drt3b that turned out to be the big surprise, doing its part of the DNA construction work without any external templates to refer to. It's an all-in-one mechanism that has never been seen before.
Our current understanding of biology suggests that genetic information flows from a DNA blueprint to the builder proteins. But with Drt3b that process is self-contained – the layout of the assembly line effectively acts as its own blueprint by dictating how the DNA is coded.
"The protein itself serves as the blueprint for the DNA sequence," Stanford biochemist Alex Gao tells Richard Stone at Science. "That was quite a surprise. This is a fundamentally new way that life produces DNA."
For now, the researchers aren't entirely sure how bacteria use DRT3 to protect themselves against attacking viruses. And while it may seem like a very specific use case, that doesn't mean we can't find other things to use it for.
CRISPR also began as a natural bacterial defense system, before scientists borrowed it to make the groundbreaking gene-editing technique.
Further down the line there's the possibility that the trick used by Drt3b could be harnessed and engineered too – though that's still a long way off.
While scientists are already working on ways to construct synthetic DNA in the lab, the particular Drt3b polymerase studied here is made as a very specific and fixed mold. It's going to be difficult, though perhaps not impossible, to reprogram it for other uses.
Related: A New DNA-Printing Technique Could Revolutionize How We Store Data
Further studies are also going to be required to understand exactly how DRT3 fights off virus attacks, and how the bacteria is using it. That should provide more insight into the construction of this kind of DNA and how it might be used.
There's also the question of how this efficient shortcut came to be. The researchers predict DRT3 will be active across many bacterial strains, and that it has a long evolutionary history as a way of combating viruses while expending as little energy as possible.
"Altogether, the DRT3 system employs an unexpected mechanism of biological information transfer, expanding the remarkable repertoire of nucleic acid–based strategies in anti-phage defense," write the researchers.
The research has been published in Science.