After decades of decline in the developed world, tuberculosis (TB) is back on the rise, and increasingly resistant to our best antibiotics.
The World Health Organization (WHO) is calling it a public health crisis – but help may be on the way to fight the world's top infectious killer.
An international team of researchers has studied three promising experimental antibiotic compounds – ecumicin, ilamycins, and cyclomarins – to determine how they take down Mycobacterium tuberculosis, the bacterium that causes TB.
While these compounds and their potential as TB treatments aren't new to scientists, there were still open questions around exactly how they might disable M. tuberculosis. That's a key detail to know along the way to developing treatments at scale.
In lab tests, the researchers were able to show that all three act on a molecular machine inside the bacterium, known by the catchy title of the ClpC1–ClpP1P2 complex. This complex is vital to M. tuberculosis, enabling it to clear out waste proteins that aren't needed or have become damaged.
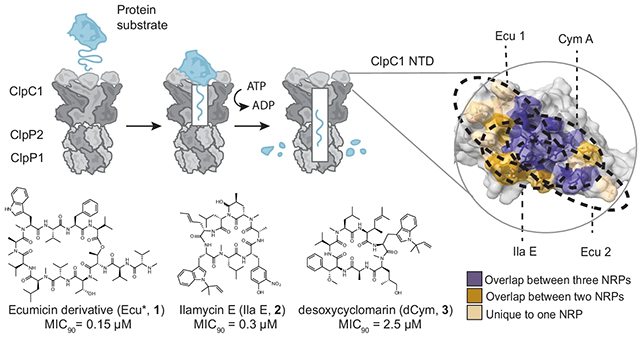
"TB bacteria depend on this recycling system to stay alive, particularly under stressful conditions inside the human body," says immunologist Warwick Britton of the University of Sydney in Australia.
"Our findings show these compounds don't simply shut the system down. Instead, each one interferes with it in a different way, triggering widespread imbalances across the whole bacterium. This disruption weakens its ability to function and survive."
The analysis techniques deployed by the researchers measured more than 3,000 proteins across M. tuberculosis. Each of the antibiotic compounds was tested in turn to see how these thousands of proteins were affected.
While all three compounds caused disruption in the bacterium's protein recycling system, they didn't all act in the same way. Ecumicin had the biggest effect out of the three, causing a protective stress protein called Hsp20 to spike (a clear sign the bacterium was under severe stress).
All of this means development of antibiotics containing these compounds can proceed with more precision – with a much clearer picture of what damage they do to M. tuberculosis and how they might be combined most effectively.
"By tracking changes across most of the bacterium's protein network, we were able to see how disrupting a single essential complex can reshape the bacterium's entire internal protein landscape," says chemical biologist Isabel Barter of the University of Sydney.
"This deeper understanding gives us valuable insight into how we might refine these compounds and design more precise and effective anti-TB treatments."
Tuberculosis is now claiming well over a million lives every year, and can be spread through airborne droplets (from coughing or breathing). While the disease is curable, effective treatments are not universally available. A full course of TB-fighting drugs can take months to fully work, which may contribute to the emergence of antibiotic-resistant strains.
Much depends on socioeconomic conditions when it comes to a person's chances of surviving TB, as well as how weak or strong their immune system is. It's also an infection that lies dormant in many people – up to a quarter of the global population are thought to have been infected with TB bacteria, though this doesn't always develop into illness.
Related: Mixing Coffee And Antibiotics Could Be a Bad Idea, Study Shows
Each step forward we take in understanding how TB takes hold in the body and how it might be countered by modern-day drugs gets us closer to eradicating it for good.
This study highlights a promising target for killing TB bacteria in their own waste proteins, and three promising compounds to target it – potentially helping us stay ahead of antibiotic-resistant strains.
"Our study highlights the potential of directly targeting this protein degradation system," says chemical biologist Richard Payne of the University of Sydney.
"By understanding how different compounds interact with it and disrupt its normal function, we can more strategically design the next-generation of anti-TB drugs."
The research has been published in Nature Communications.