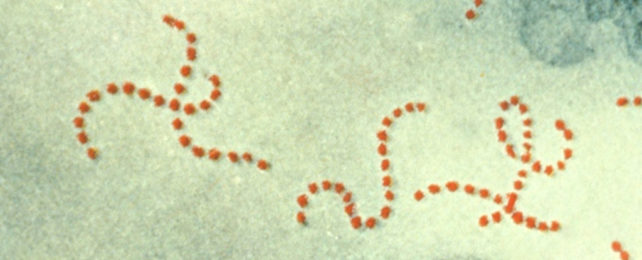
Researchers have just caught bacteria sidestepping antibiotic treatment with a never-before-seen trick.
Bacteria's troublesome talent for developing resistance against antibiotics is a rapidly growing health threat. This ability has ancient origins and allows drug-resistant bacterial infections like MRSA and gonorrhea to kill 1.3 million people globally each year.
These superbugs are even finding their way into wild animals, such as dolphins and bears.
The shifty microbes can steal genes from each other, quickly passing on antibiotic-resistant tactics: Strategies include directly inactivating the antibiotics, preventing antibiotics from accumulating in their systems, or changing the antibiotic's targets so that the drugs are no longer effective.
Thanks in part to antibiotic overuse, superbugs have been accumulating multiple resistant tactics, making them extremely difficult to treat.
"This new form of resistance is undetectable under conditions routinely used in pathology laboratories, making it very hard for clinicians to prescribe antibiotics that will effectively treat the infection, potentially leading to very poor outcomes and even premature death," explains Telethon Kids Institute infectious disease researcher Timothy Barnett.
Telethon Kids Institute microbiologist Kalindu Rodrigo and colleagues discovered this new mechanism while investigating how Group A Streptococcus responds to antibiotics.
This bacteria commonly cause sore throats and skin infections, but it can also lead to systemic infections like scarlet fever and toxic shock syndrome.
"Bacteria need to make their own folates to grow and, in turn, cause disease. Some antibiotics work by blocking this folate production to stop bacteria growing and treat the infection," explains Barnett.
"When looking at an antibiotic commonly prescribed to treat Group A Strep skin infections, we found a mechanism of resistance where, for the first time ever, the bacteria demonstrated the ability to take folates directly from its human host when blocked from producing their own."
So Streptococcus has been acquiring already processed folate from outside its own cells; these molecules are abundant in our bodies.
The process completely bypasses the action of sulfamethoxazole, an antibiotic that inhibits folate synthesis within the bacteria, thus rendering the drug ineffective.
Rodrigo and the team identified at least one gene involved: thfT. It encodes part of the folate harvesting system, not unlike our own, as we also can't produce folate and must get it from our food.
Streptococcus bacteria with this gene, therefore, have found a way to suck up folate and subvert sulfamethoxazole.
In the lab, Group A Streptococcus does succumb to sulfamethoxazole antibiotics because it doesn't have another accessible source of folate.
In this case, the bacteria are only resistant to the antibiotics when they're causing an actual infection inside our bodies. This means there's no easy way of detecting this antibiotic resistance in pathology labs yet.
This mechanism suggests antibiotic resistance is far more varied than researchers previously realized and emphasizes the need to establish more diverse treatments against bacteria.

"Unfortunately, we suspect this is just the tip of the iceberg – we have identified this mechanism in Group A Strep, but it's likely it will be a broader issue across other bacterial pathogens," says Barnett.
Understanding these mechanisms is the first step towards being able to test for them and counter them by prescribing other classes of antibiotics instead.
"It is vital we stay one step ahead of the challenges of antimicrobial resistance and, as researchers, we should continue to explore how resistance develops in pathogens and design rapid accurate diagnostic methods and therapeutics," urges Rodrigo.
This research was published in Nature Communications.